Using boost-histogram#
[1]:
import functools
import operator
import matplotlib.pyplot as plt
import numpy as np
import boost_histogram as bh
1: Basic 1D histogram#
Let’s start with the basics. We will create a histogram using boost-histogram and fill it.
1.1: Data#
Let’s make a 1d dataset to run on.
[2]:
data1 = np.random.normal(3.5, 2.5, size=1_000_000)
Now, let’s make a histogram
[3]:
hist1 = bh.Histogram(bh.axis.Regular(40, -2, 10))
[4]:
hist1.fill(data1)
[4]:
Histogram(Regular(40, -2, 10), storage=Double()) # Sum: 981542.0 (1000000.0 with flow)
You can see that the histogram has been filled. Let’s explicitly check to see how many entries are in the histogram:
[5]:
hist1.sum()
[5]:
981542.0
What happened to the missing items? They are in the underflow and overflow bins:
[6]:
hist1.sum(flow=True)
[6]:
1000000.0
Like ROOT, we have overflow bins by default. We can turn them off, but they enable some powerful things like projections.
Let’s plot this (Hist should make this easier):
[7]:
plt.bar(hist1.axes[0].centers, hist1.values(), width=hist1.axes[0].widths);
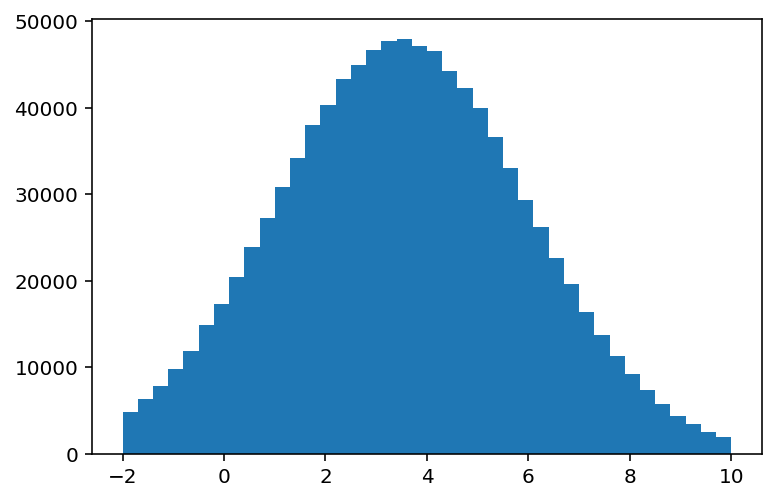
Note: you can select the axes before or after calling .centers; this is very useful for ND histograms.
From now on, let’s be lazy
[8]:
plothist = lambda h: plt.bar(*h.axes.centers, h.values(), width=h.axes.widths[0]);
Aside: here’s step. The edges are quite ugly for us, just like it is for numpy. Or anyone.
[9]:
plt.step(hist1.axes[0].edges[:-1], hist1.values(), where="post");
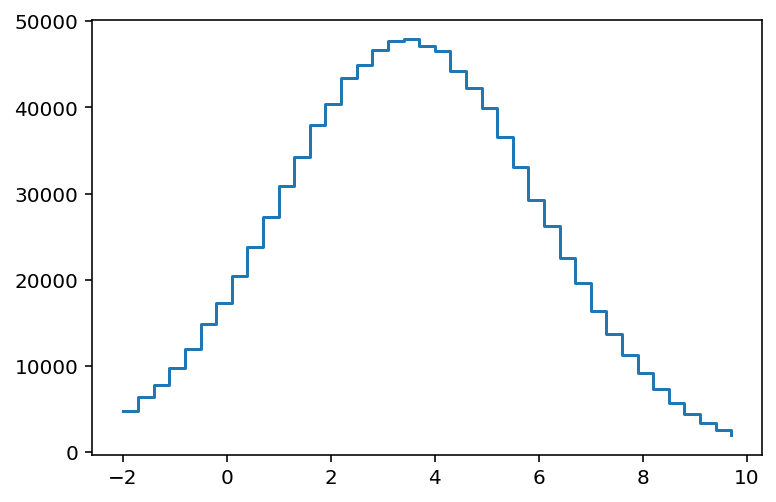
Recent versions of matplotlib support .stairs, which was designed to work well with histograms:
[10]:
plt.stairs(hist1.values(), hist1.axes[0].edges);
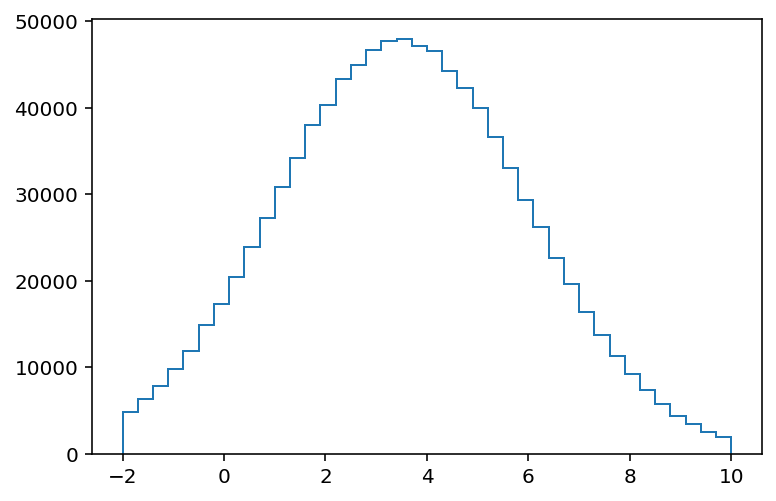
No plotting is built in, but the data is easy to access.
2: Drop-in replacement for NumPy#
To start using this yourself, you don’t even need to change your code. Let’s try the numpy adapters.
[11]:
bins2, edges2 = bh.numpy.histogram(data1, bins=10)
[12]:
b2, e2 = np.histogram(data1, bins=10)
[13]:
bins2 - b2
[13]:
array([0, 0, 0, 0, 0, 0, 0, 0, 0, 0], dtype=int64)
[14]:
e2 - edges2
[14]:
array([ 0.00000000e+00, 8.88178420e-16, 8.88178420e-16, 0.00000000e+00,
0.00000000e+00, 0.00000000e+00, 1.77635684e-15, 3.55271368e-15,
0.00000000e+00, 0.00000000e+00, -1.77635684e-15])
Not bad! Let’s start moving to the boost-histogram API, so we can use our little plotting function:
[15]:
hist2 = bh.numpy.histogram(data1, bins="auto", histogram=bh.Histogram)
plothist(hist2);
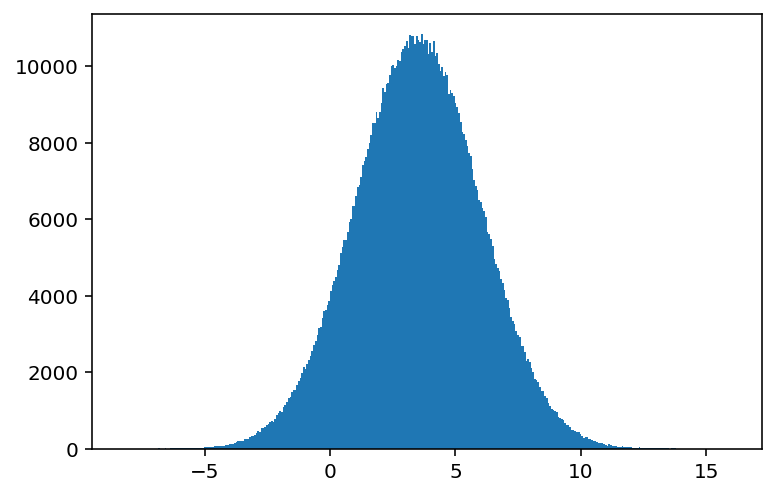
Now we can move over to boost-histogram one step at a time! Just to be complete, we can also go back to a NumPy tuple from a Histogram object:
[16]:
b2p, e2p = bh.numpy.histogram(data1, bins=10, histogram=bh.Histogram).to_numpy()
b2p == b2
[16]:
array([ True, True, True, True, True, True, True, True, True,
True])
3: More dimensions#
The same API works for multiple dimensions.
[17]:
hist3 = bh.Histogram(bh.axis.Regular(150, -1.5, 1.5), bh.axis.Regular(100, -1, 1))
[18]:
def make_2D_data(*, mean=(0, 0), widths=(1, 1), size=1_000_000):
cov = np.asarray(widths) * np.eye(2)
return np.random.multivariate_normal(mean, cov, size=size).T
[19]:
data3x = make_2D_data(mean=[-0.75, 0.5], widths=[0.2, 0.02])
data3y = make_2D_data(mean=[0.75, 0.5], widths=[0.2, 0.02])
From here on out, I will be using .reset() before a .fill(), just to make sure each cell in the notebook can be rerun.
[20]:
hist3.reset()
hist3.fill(*data3x)
hist3.fill(*data3y)
[20]:
Histogram(
Regular(150, -1.5, 1.5),
Regular(100, -1, 1),
storage=Double()) # Sum: 1905785.0 (2000000.0 with flow)
Again, let’s make plotting a little function:
[21]:
def plothist2d(h):
return plt.pcolormesh(*h.axes.edges.T, h.values().T)
This is transposed because pcolormesh expects it.
[22]:
plothist2d(hist3);
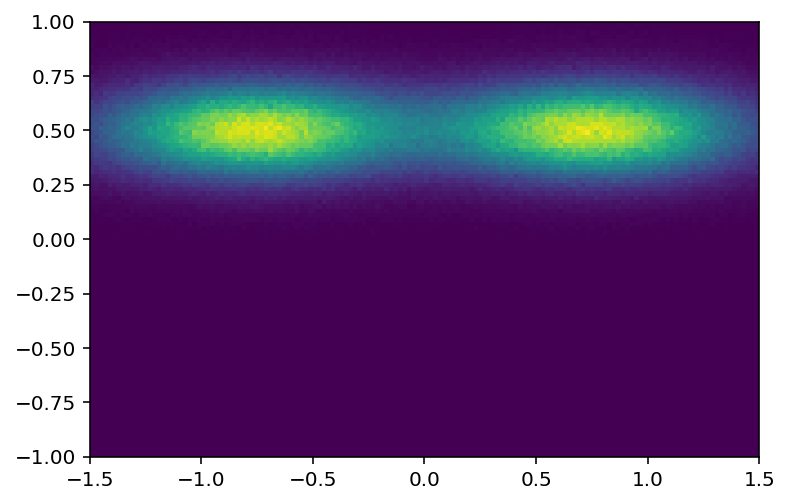
Let’s try a 3D histogram
[23]:
data3d = [np.random.normal(size=1_000_000) for _ in range(3)]
hist3d = bh.Histogram(
bh.axis.Regular(150, -5, 5),
bh.axis.Regular(100, -5, 5),
bh.axis.Regular(100, -5, 5),
)
hist3d.fill(*data3d)
[23]:
Histogram(
Regular(150, -5, 5),
Regular(100, -5, 5),
Regular(100, -5, 5),
storage=Double()) # Sum: 1000000.0
Let’s project to the first two axes:
[24]:
plothist2d(hist3d.project(0, 1));
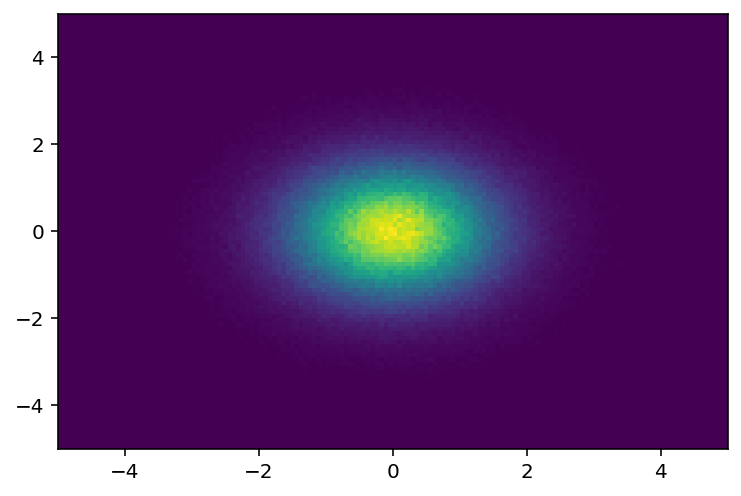
4: UHI#
Let’s explore the boost-histogram UHI syntax. We will reuse the previous 2D histogram from part 3:
[25]:
plothist2d(hist3);
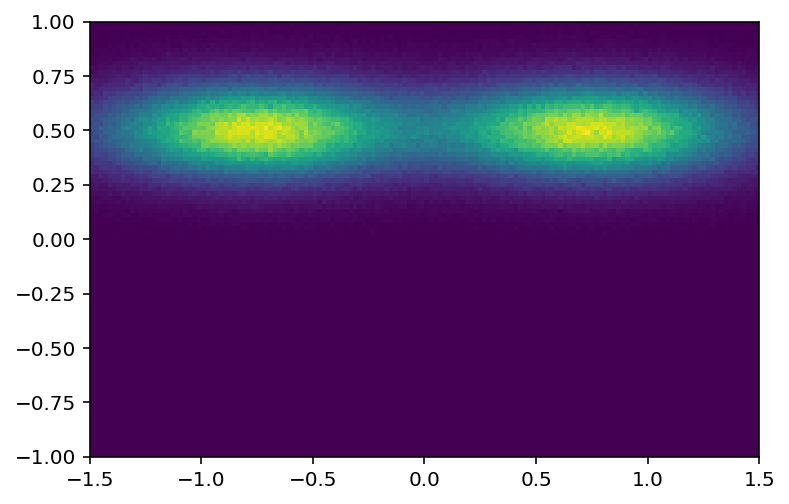
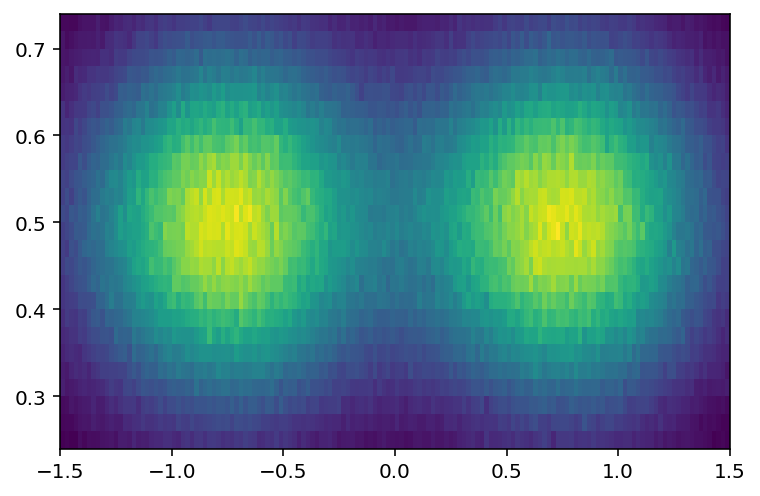
I can see that I want y from 0.25 to 0.75, in data coordinates:
[26]:
plothist2d(hist3[:, bh.loc(0.25) : bh.loc(0.75)]);
What’s the contents of a bin?
[27]:
hist3[100, 87]
[27]:
181.0
How about in data coordinates?
[28]:
hist3[bh.loc(0.5), bh.loc(0.75)]
[28]:
181.0
Note: to get the coordinates manually:
hist3.axes[0].index(.5) == 100 hist3.axes[1].index(.75) == 87
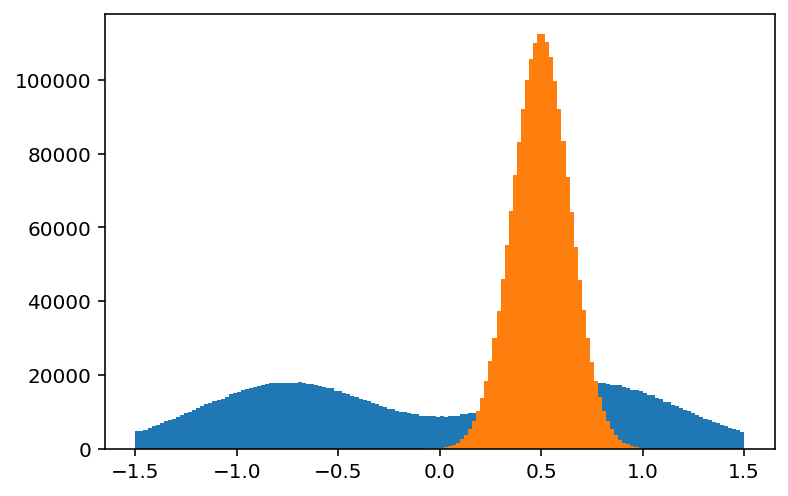
How about a 1d histogram?
[29]:
plothist(hist3[:, :: bh.sum])
plothist(hist3[:: bh.sum, :]);
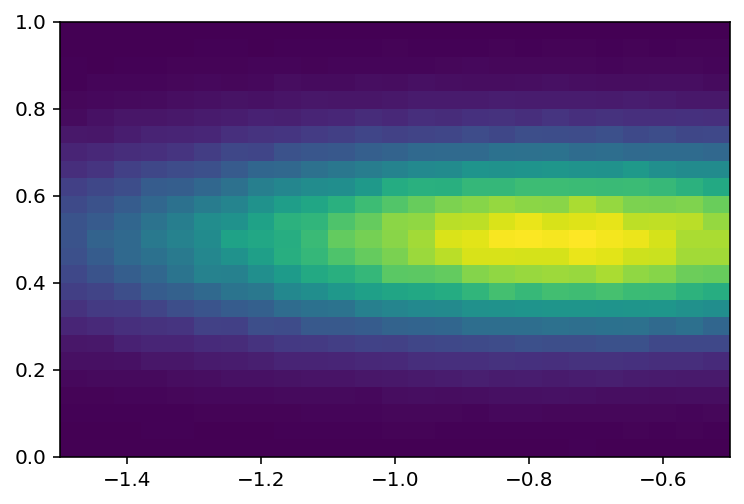
Let’s look at one part and rebin:
[30]:
plothist2d(hist3[: 50 : bh.rebin(2), 50 :: bh.rebin(2)]);
What is the value at (-.75, .5)?
[31]:
hist3[bh.loc(-0.75), bh.loc(0.5)]
[31]:
1005.0
5: Understanding accumulators#
Boost-histogram has several different storages; storages store accumulators. Let’s try making a profile.
[32]:
mean = bh.accumulators.Mean()
mean.fill([0.3, 0.4, 0.5])
[32]:
Mean(count=3, value=0.4, variance=0.01)
Here’s a quick example accessing the values:
[33]:
print(f"{mean.count=} {mean.value=} {mean.variance=}")
mean.count=3.0 mean.value=0.4 mean.variance=0.01
6: Changing the storage#
[34]:
hist6 = bh.Histogram(bh.axis.Regular(10, 0, 10), storage=bh.storage.Mean())
[35]:
hist6.fill([0.5] * 3, sample=[0.3, 0.4, 0.5])
[35]:
Histogram(Regular(10, 0, 10), storage=Mean()) # Sum: Mean(count=3, value=0.4, variance=0.01)
[36]:
hist6[0]
[36]:
Mean(count=3, value=0.4, variance=0.01)
[37]:
hist6.view()
[37]:
MeanView(
[(3., 0.4, 0.02), (0., 0. , 0. ), (0., 0. , 0. ), (0., 0. , 0. ),
(0., 0. , 0. ), (0., 0. , 0. ), (0., 0. , 0. ), (0., 0. , 0. ),
(0., 0. , 0. ), (0., 0. , 0. )],
dtype=[('count', '<f8'), ('value', '<f8'), ('_sum_of_deltas_squared', '<f8')])
[38]:
hist6.view().value
[38]:
array([0.4, 0. , 0. , 0. , 0. , 0. , 0. , 0. , 0. , 0. ])
[39]:
hist6.view().variance
[39]:
array([ 0.01, -0. , -0. , -0. , -0. , -0. , -0. , -0. , -0. ,
-0. ])
7: Making a density histogram#
Let’s try to make a density histogram like NumPy’s.
[40]:
bins = [
-10,
-7,
-4,
-3,
-2,
-1,
-0.75,
-0.5,
-0.25,
0,
0.25,
0.5,
0.75,
1,
2,
3,
4,
7,
10,
]
d7, e7 = np.histogram(data1 - 3.5, bins=bins, density=True)
plt.hist(data1 - 3.5, bins=bins, density=True);
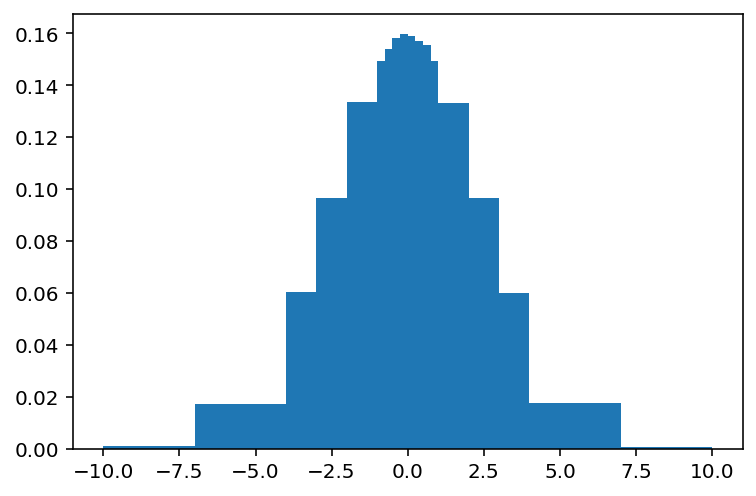

Yes, it’s ugly. Don’t judge.
We don’t have a .density! What do we do? (note: density=True is supported if you do not return a bh object)
[41]:
hist7 = bh.numpy.histogram(data1 - 3.5, bins=bins, histogram=bh.Histogram)
widths = hist7.axes.widths
area = functools.reduce(operator.mul, hist7.axes.widths)
area
[41]:
array([3. , 3. , 1. , 1. , 1. , 0.25, 0.25, 0.25, 0.25, 0.25, 0.25,
0.25, 0.25, 1. , 1. , 1. , 3. , 3. ])
Yes, that does not need to be so complicated for 1D, but it’s general.
[42]:
factor = np.sum(hist7.values())
view = hist7.values() / (factor * area)
[43]:
plt.bar(hist7.axes[0].centers, view, width=hist7.axes[0].widths);
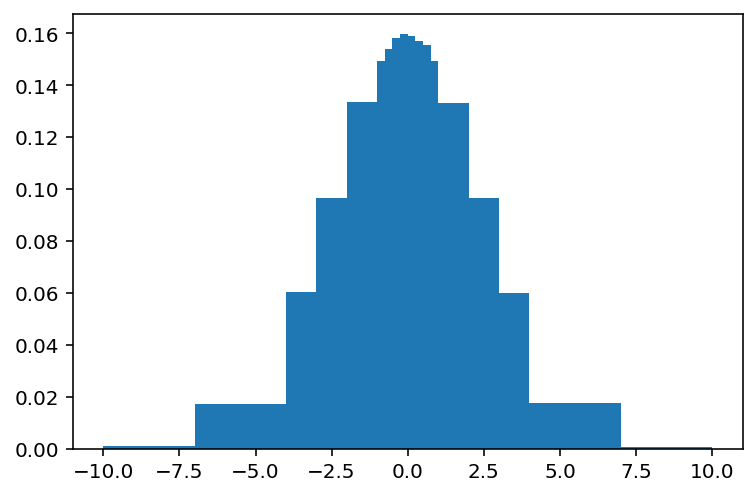

8: Axis types#
There are more axes types, and they all provide the same API in histograms, so they all just work without changes:
[44]:
hist8 = bh.Histogram(
bh.axis.Regular(30, 1, 10, transform=bh.axis.transform.log),
bh.axis.Regular(30, 1, 10, transform=bh.axis.transform.sqrt),
)
[45]:
hist8.reset()
hist8.fill(*make_2D_data(mean=(5, 5), widths=(5, 5)))
[45]:
Histogram(
Regular(30, 1, 10, transform=log),
Regular(30, 1, 10, transform=sqrt),
storage=Double()) # Sum: 903807.0 (1000000.0 with flow)
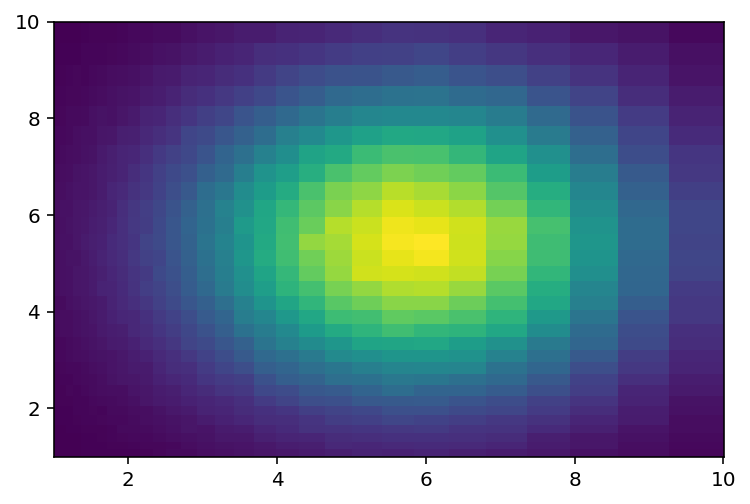
[46]:
plothist2d(hist8);
9: And, circular, too!#
[47]:
hist9 = bh.Histogram(bh.axis.Regular(30, 0, 2 * np.pi, circular=True))
hist9.fill(np.random.uniform(0, np.pi * 4, size=300))
[47]:
Histogram(Regular(30, 0, 6.28319, circular=True), storage=Double()) # Sum: 300.0
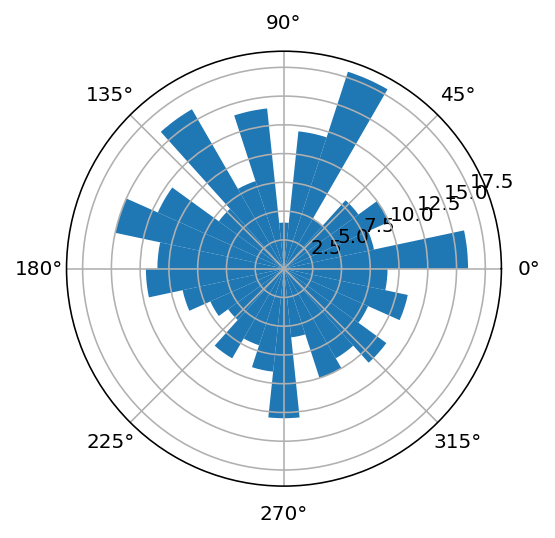
Now, the really complicated part, making a circular histogram:
[48]:
ax = plt.subplot(111, polar=True)
plothist(hist9);