ROOT file format example#
To run this example, you will need uproot, which is another SciKit-HEP library.
[1]:
from pathlib import Path
import matplotlib.pyplot as plt
import numpy as np
import uproot
import boost_histogram as bh
demo_file = Path("demo_uproot_file.root")
ROOT is a modular scientific software toolkit used in High Energy Physics. (HEP) The ROOT file format is used to store almost all HEP data. This notebook will illustrate one method for converting to/from the ROOT file format using uproot, a Python implementation of a ROOT file reader and writer.
For more complicated histograms, you may need Aghast and PyROOT, but that is a much heaver dependency, and is covered in a separate tutorial.
Start by making a 1D histogram:
[2]:
h = bh.Histogram(bh.axis.Regular(15, -3, 3))
h.fill(np.random.normal(size=1_000_000))
[2]:
Histogram(Regular(15, -3, 3), storage=Double()) # Sum: 997352.0 (1000000.0 with flow)
[3]:
with uproot.recreate(demo_file) as root_file:
# Uproot automatically converts histograms
root_file["hist"] = h
If you want to save and load the ROOT histogram, use uproot to read and write:
[4]:
with uproot.open(demo_file) as root_file_2:
uproot_hist = root_file_2["hist"]
print(uproot_hist)
<TH1D (version 3) at 0x7fa9b83ffd00>
This uproot histogram can be converted directly to boost_histogram:
[5]:
h = bh.Histogram(uproot_hist)
plt.bar(h.axes[0].centers, h.values(), width=h.axes[0].widths);
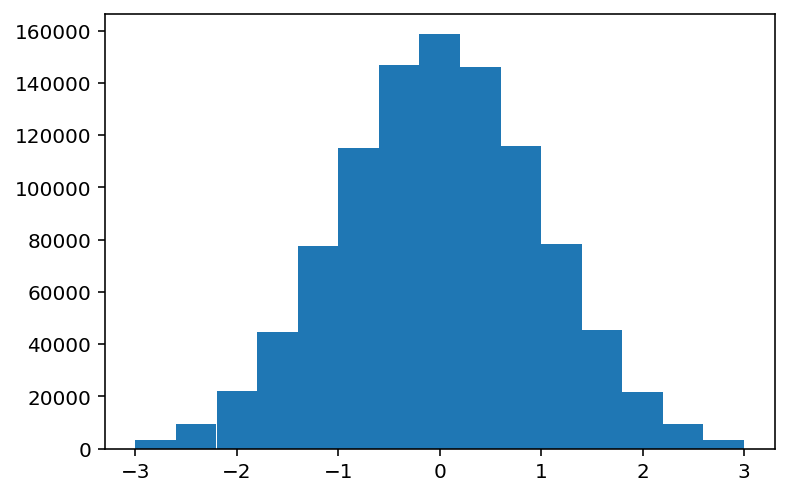
We could use a Weight() storage and read both allvalues and allvariances in, as well, since ROOT histograms can sometimes have this enabled.
Finally, let’s clean up after ourselves:
[6]:
if demo_file.is_file():
demo_file.unlink()
